Horizontal gene transfer/Citable Version
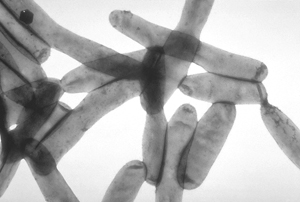
Horizontal gene transfer (HGT; also called lateral gene transfer, LGT) is any process in which an organism transfers genetic material to another cell or organism that is not one of its own offspring. HGT is thus very different from the normal vertical gene transfer whereby parental traits are inherited by progeny by sexual fusion of gametes to form zygotes as in animals and plants, or by asexual reproduction as in microorganisms such as bacteria and fungi. HGT occurs at a much lower frequency than vertical gene transfer, so is not easily detected directly, and special techniques are needed to document it.
Introduction
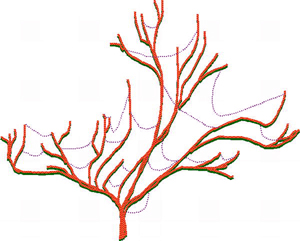
Advances in genome science and bioinformatics have brought abundant evidence that natural HGT has occurred between diverse biological taxa that are widely separated in the phylogenetic tree of life. Horizontal gene transfers include gene movement between different species of microbes and other microbial taxa such as protists, gene movement between different plant families, between different animals, and between bacteria and plants. Microorganisms are most affected by HGT, but recent research suggests that HGT in microbes is limited to about 2% of core genes in microbial genomes, and this is low enough to allow the the main lineages of microbial evolution to be treated as 'trees' with HGT 'cobwebs' (see figure to right).
Gene transfers between different biological sub-kingdoms (domains), such as between eukaryotic protists and bacteria [1] , or between bacteria and insects [2] are the most phylogenetically extreme cases of HGT. An example is bacterial "rol" genes from Agrobacterium species which have been found in tobacco plants (Nicotiniana). [3].
HGT is just one of the processes causing rearrangement of genomes during evolution. The possibility of intracellular movement of genes between different parts of an organism's genome (that is, between the chromosomes of the nucleus, the circular mitochondrion chromosome , and the circular plastid (chloroplast) chromosome) need to be considered when evaluating HGT between different species.[4]
Main features of HGT in nature
- A hallmark of HGT is the presence of the same gene in organisms that are only very distantly related. The frequent discovery of shared DNA sequences such as the mariner class of transposons, insertion sequence DNA, and retrovirus genes in diverse species, and shared mitochondrial genes in diverse flowering plants, indicate that mobile DNA has natural pathways for movement between species. (The name mariner for a class of related transposons an allusion to The Rhyme of the Ancient Mariner, meaning a traveller to distant lands.) Close relatives of mariner mobile DNA have been discovered in organisms as diverse as mites, flatworms, hydras, insects, nematodes, mammals and humans.[5]

- Horizontal movement of genes is common among bacteria, and is a major factor in accelerating the rate of evolution in these organisms.[6] HGT is involved in infectious multiple-antibiotic resistance in pathogenic bacteria, a major factor limiting the effectiveness of antibiotics. Inter-domain(sub-kingdom) transfer of several genes from eukaryotes to bacteria for instance, has occurred in the "accidentally pathogenic" bacterium (Legionella pneumophila, see illustration) that lives and replicates within a vacuoles of protist and mammalian macrophage cells.[7]
- HGT is common in diverse groups of unicellular protists, which often contain several genes transferred from both prokaryotes and other protists. [8]
- HGT occurs globally on a massive scale among marine microorganisms, and viruses, at total numbers near 1029 being the most common biological entities in the sea, are a major pathway for inter-species gene movement in the ocean. The estimated virus-mediated gene transfer events in the Mediterranean sea are 1013 per year.[9] Endosymbiosis with an alga is identified as a route for HGT in marine dinoflagellates, the organisms that cause 'red tides'.[10]
- Interspecies gene movement within the same biological genus by cross-hybridization is common in flowering plants.[11] Mechanisms for HGT in flowering plants between more distant taxa involving parasitic plants such as dodder and endophytes such as mosses (which facilitate inter-species gene transfer by being in intimate cell-to-cell contact with their host plants) are also well established (see Horizontal gene transfer in plants). Plant mitochondria are unusually active in HGT. [12]
- Not all of the vehicles by which HGT occurs are fully characterized, but some are clearly identified. HGT is difficult to detect directly, as it occurs at lower frequencies than with normal sexual reproduction within the species, but modern techniques of DNA analysis, by providing detailed comparison of genomes, provide much of the evidence for HGT. In insects, mites and viruses are probable vectors for HGT. In bacteria, surface appendages called pili have various roles in DNA uptake, DNA secretion and DNA transfer which have been extensively analyzed; HGT in bacteria includes plasmid-mediated promiscuous mating by bacteria, for instance by the crown-gall bacterium Agrobacterium tumefaciens[13], and carriage of genes between species by viruses.[14] Direct DNA uptake as another transfer mechanism is illustrated by Legionella bacteria, which are naturally competent for DNA uptake.
Prokaryotes
- See main article HGT in prokaryotes
- The three main mechanisms of HGT in bacteria and archaea discussed here are:
- Bacterial Transformation or direct uptake of extracellular DNA.
- Transduction of genes by bacterial viruses.
- Bacterial conjugation, a gene transfer process carried out by plasmids and conjugative transposons.
Eukaryotes
- See also Endosymbiotic theory
Protists
Analysis of the complete genome sequence of the protist Entamoeba histolytica indicates 96 cases of relatively recent HGT from prokaryotes [15], whereas similar analysis of the complete genome sequence of the protist Cryptosporidium parvum reveals 24 candidates of HGT from bacteria [16].There is also convincing evidence that a bacterial gene for a biosynthetic enzyme has been recruited by the protist Trichomonas vaginalis from bacteria related to the ancestors of Pasteurella bacteria.[17] These results fit the idea that "you are what you eat". That is, with unicellular grazing organisms, foreign genetic material is constantly entering the cell and occasionally the genome from food organisms [18]
Fungi
Comparison of the genome sequences of two fungi, baker's yeast (Saccharomyces cerevisiae) and Ashbya gossypii, has shown that Saccharomyces has received two genes from bacteria by HGT. One codes for an enzyme that allows baker's yeast to make pyrimidine nucleotide bases anaerobically, and the other allows usage of sulfur from several organic sulfur sources.[19]. Other work with yeasts suggests that eight genes from Yarrowia lipolytica, five from Kluyveromyces lactis, and one from Debaryomyces hansenii are horizontally transferred. [20]
Other eukaryotes
Analysis of DNA sequences suggests that HGT has also occurred within multicellular eukaryotes, by a route that involves transfer of genes from chloroplast and mitochondrial genomes to the nuclear genomes. [21] According to the endosymbiotic theory, chloroplasts and mitochondria originated as the bacterial endosymbionts of a progenitor to the eukaryotic cell, and endosymbiosis can be considered to be a special case of HGT.
Plants
- See Horizontal gene transfer in plants for
- Natural gene transfer between plants that do not cross-pollinate
- Jumping genes cross naturally between rice and millet
- Epiphytes and parasites as a bridge for gene flow between diverse plant species
- See Transgenic plant for hybridization by cross-pollination and artificial horizontal gene transfer in biotechnology.
Plant genes have also been discovered to be able to move to endophyte fungi that grow on them. Several plant endophyte fungi that grow on taxol-producing yew trees have gained the ability to make taxol themselves [22]. (Taxol, also called paclitaxel, is an anti-cancer drug found in yew trees.)
Animals
Junk DNA is the most obvious general evidence of HGT in eukaryotes. Such seemingly non-functional repetitive DNA is a major portion of many genomes of plants and animals. This DNA usually includes multiple copies of various "Jumping genes" which can proliferate within a genome after they have been transferred from another species. Examples in the human of such mobile elements are 'Hsmar1' and 'Hsmar2' which are related to the widely studied 'mariner' transposon. Close relatives of mariner mobile DNA have been discovered in organisms as diverse as mites, flatworms, hydras, insects, nematodes, mammals and humans [23].Retroviruses and retrotransposons are other examples of mobile horizontally transferred DNA found in animals.
The adzuki bean beetle Callosobruchus chinensis is infected with several strains of bacterial Wolbachia endosymbionts. A genome fragment of one of these endosymbionts has been found transferred to the X chromosome of the host insect [24].
History of discovery of HGT
- See main article Horizontal gene transfer (History)
- Bacterial genetics starts in 1946
- see main article Horizontal gene transfer in prokaryotes
- First glimpses of horizontal transfer of traits in plant evolution
- see also main article Barbara McClintock
- Discovery of mobile genes in flies, and mariner
- HGT and genetic engineering
Decoding the tree of life from genomes scrambled by HGT
Because each organism carries a record of its ancestry in its DNA, methods for rapid gene isolation and analysis of DNA encoded sequences - which can extract the information from this ancestral archive - have been powerful tools for answering questions about evolution. These techniques have enabled rapid expansion of the field of biological systematics known as phylogenetic inference. Comparison of related (homologous) gene sequences from different organisms has been successfully applied to reconstruct the history of many evolutionary lineages. [25]
Many issues about evolution have been clarifed by comparing homologous genes from different species, genera, families, and phyla. Unfortunately, HGT complicates the use of gene sequence evidence to reconstruct evolutionary history because HGT scrambles the evidence needed to deduce the branching patterns of evolutionary trees, and requires coalescence of branches in tree representations of evolution. One area of current research in phylogenetic inference, and arguably one the most challenging problems in evolutionary theory, is the early stages in the evolution of life. This quandary about the origins of different cell types provides a good illustration of the complications introduced by HGT into the reconstruction of evolutionary history.
The main early branches of the tree of life have been intensively studied by microbiologists because the first organisms were microorganisms. A gene very commonly used for constructing phylogenetic relationships in microrganisms is the small ribosomal subunit ribosomal RNA gene (SSU rRNA) gene, as its sequences tend to be conserved among members with close phylogenetic distances, yet it is variable enough that differences can be measured [26]. The SSU rRNA as a measure of evolutionary distances was pioneered by Carl Woese when formulating the first modern "tree of life", and his results led him to propose the Archaea as a third domain (sub-kingdom) of life.
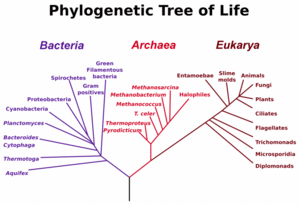
Microbiologists introduced the term domain for the three main early branches of the tree of life, where domain is a phylogenetic term very similar in meaning to biological kingdom. These represent the three main lineages in evolution of early cellular life, and are currently represented by the Bacteria, the Archaea (single celled organisms superficially similar to bacteria), and Eukarya (eukaryote) domains. Eukaryotes are all organisms with a well defined nucleus, and this domain comprises protists, fungi, and all organisms in the animal and plant kingdoms, including humans. As seen in the figure (left), studies of SSU rRNA genes (and some other genes) might suggest that Archaea and Eukarya have a sister relationship in evolution, and it has often been assumed that eukaryotes evolved from archeal cells. However, the fact that genes can move between distant branches of the tree of life even at low probabilities poses problems for scientists trying to reconstruct evolution from studying genes and gene sequences in different organisms due to the scrambling effect of HGT. The challenges are most awkward for the ambitious reconstruction of the earliest branches of the tree of life - because over a long enough time and with large numbers of organisms, many HGT events are certain to have occurred even though each particular transfer event has a low probability.
Thus recent discoveries of 'rampant' HGT in microorganisms, and the detection of horizontal movement of even genes for SSU rRNA have forced biologists to question the accuracy of at least the early branches in the tree of life, and even question the validity of trees as useful models of microbial evolution.[27]
Recent efforts to infer evolutionary trees while recognizing HGT
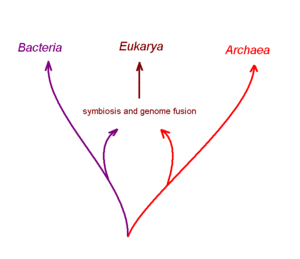
The challenges of acertaining an accurate branching structure for evolutionary trees of microbes, especially the earliest branches in the tree of life, have been a great spur for current biological research. Clear answers to important questions about these trees are not yet available, but many interesting discoveries and ideas are emerging.
Instead of relying a single gene such as the SSU rRNA gene to reconstruct evolution, scientific effort has shifted to exploiting the comprehensive information from the many complete genome sequences of organisms that are now available.[28]
So far, this comparitive genomics approach, made possible by specially developed computer programs and mathematical algorithms, suggests that most of the core genes of bacteria that are useful for deducing evolutionary histories are mainly unaffected by HGT. This indicates that reliable trees can still be deduced for a least the more recent stages in microbial evolution, such as evolutionary relationships within particular bacterial phyla, provided that many genes are used to investigate how lineages are related to one another. Thus it is argued that the 'tree' is still a valid metaphor for microbial evolution - but a tree adorned with 'cobwebs' of horizontally transferred genes.
But it is also clear that trees based only on SSU rRNA alone do not capture the events of early eukaryote evolution accurately, and the origins of the first nucleated cells are still uncertain. For instance, careful analysis of the complete genome of the eukaryote yeast shows that it contains many genes more that closely related to bacterial genes than they are to archaea, and it is now clear that archaea were not the simple progenitors of the eukaryotes, in stark contradiction to the findings based on SSU rRNA and limited samples of other genes. [29]
One controversial hypothesis coming out recent computer assisted studies of complete genome DNA sequences is that the first nucleated cell arose from two distinctly different ancient prokaryotic (non-nucleated) species that had formed a symbiotic relationship with one-another to carry out different aspects of metablism. One partner of this postulated symbiosis is proposed to be a true bacterial cell, and the other an archaean cell. It is postulated that this symbiotic partnership progressed to the cellular fusion of the partners to generate a chimeric or hybrid cell with a membrane bound internal structure that was the forerunner of the nucleus. The next stage in this scheme was transfer of both partner genomes into the nucleus and their fusion with one-another. Several variations on this chimera hypothesis for the origin of nucleated cells have been suggested[30], but other biologists dispute the proposal.[31]
See also
- Gene flow
- Species
- Phylogenetic tree
- Endogenous retrovirus
- Germline
- Mitochondrion
- Endosymbiotic theory
- Integron
- Provirus
- Retrotransposon
- Mobile DNA
- Pilus
- Transgenic plant
References
Citations
- ↑ de Felipe K et al. (2005) Evidence for acquisition of Legionella type IV secretion substrates via interdomain horizontal gene transfer. J Bacteriol 187:7716-26 PMID 16267296
- ↑ Kondo N et al. (2002) Genome fragment of Wolbachia endosymbiont transferred to X chromosome of host insect. Proc Natl Acad Sci USA 99:14280-5 PMID 12386340
- ↑ Intrieri MC, Buiatti M (2001) The horizontal transfer of Agrobacterium rhizogenes genes and the evolution of the genus Nicotiana. Mol Phylogen Evol 20:100-10 PMID 11421651
- ↑ Adams KL Palmer JD (2003) Evolution of mitochondrial gene content: gene loss and transfer to the nucleus. Molecular Phylogenetics and Evolution 29:380–95. PMID 14615181
- Timmis JN et al. (2004) Endosymbiotic gene transfer: organelle genomes forge eukaryotic chromosomes. Nat Rev Genet 5:123-35. PMID 14735123
- ↑ Robertson HM (1993) The mariner transposable element is widespread in insects. Nature 362:241-5 PMID 8384700
- Robertson HM (1996) Reconstruction of the ancient mariners of humans. Nature Genetics 12:360-1 PMID 8630486
- ↑ Jain R et al. (2003) Horizontal gene transfer accelerates genome innovation and evolution. Mol Biol Evol 20:1598-602 PMID 12777514
- (2005) Where Do All Those Genes Come From? PLoS Biol 3(5): e169 doi:10.1371/journal.pbio.0030169
- ↑ de Felipe KS et al. (2005) Evidence for acquisition of Legionella type IV secretion substrates via interdomain horizontal gene transfer. J Bacteriol 187:7716-26 PMID 16267296
- ↑ Richards TA et al. (2003) Protist 1:17–32 PMID 12812367
- Coombs GH et al. (2004) The amitochondriate eukaryote Trichomonas vaginalis contains a divergent thioredoxin-linked peroxiredoxin antioxidant system. J Biol Chem. 279:5249-56. PMID 14630923
- Andersson JO et al. (2006) Evolution of four gene families with patchy phylogenetic distributions: influx of genes into protist genomes. BMC Evol Biol PMID 16551352
- de Koning AP et al. (2000) Lateral gene transfer and metabolic adaptation in the human parasite Trichomonas vaginalis. Mol Biol Evol PMID 11070064
- Loftus B et al. (2005) The genome of the protist parasite Entamoeba histolytica. Nature 433:865-8 PMID 15729342
- Huang J et al. (2004) Phylogenomic evidence supports past endosymbiosis, intracellular and horizontal gene transfer in Cryptosporidium parvum. Genome Biology 5:R88
- ↑ Weinbauer MG et al. (2004) Are viruses driving microbial diversification and diversity? Envir Microbiol 6:1-11 PMID 14686936
- Paul JH (1999) Microbial gene transfer. J Mol Microbiol Biotechnol 1:45–50 PMID 10941783
- Fuhrman JA (1999) Marine viruses and their biogeochemical and ecological effects Nature 399:541–8 PMID 10376593
- ↑ Yoon HS et al. (2005) Tertiary endosymbiosis driven genome evolution in dinoflagellate algae. Mol Biol Evol 22:1299-308 PMID 15746017
- ↑ Sang T et al. (1995) Documentation of reticulate evolution in peonies (Paeonia) using internal transcribed spacer sequences of nuclear ribosomal DNA implications for biogeography and concerted evolution. Proc Natl Acad Sci. USA 92:6813–17 PMID 7624325
- ↑ Richardson AO, Palmer, JD (2007) Horizontal gene transfer in plants. Journal of Experimental Botany 58:1–9 doi:10.1093/jxb/erl148 PMID 17030541
- ↑ Zhu J et al. (2000) The bases of crown gall tumorigenesis. J Bacteriol 182:3885-95 PMID 10869063 This article describes the biology of crown-gall bacterium, and the mechanism of DNA injection by this bacterium, and explains how genes can move between bacterial species and from bacteria to eukaryotic organisms, and illustrates the extent to which different species can co-evolve
- ↑ Weinbauer et al. (2004) Are viruses driving microbial diversification and diversity? Envir Microbiol 6:1-11PMID 14686936
- Amoils S (2005) Analysing incompatibility — Wolbachia on the couch. Nature Rev Microbiol 3:667
- Besser TE et al. (2006) Greater diversity of Shiga toxin-encoding bacteriophage insertion sites among Escherichia coli O157:H7 isolates from cattle than from humans. Appl Environ Microbiol PMID 17142358
- ↑ Loftus B et al. (2005) The genome of the protist parasite Entamoeba histolytica. Nature 433:865-8 PMID 15729342
- ↑ Huang J et al. (2004) Phylogenomic evidence supports past endosymbiosis, intracellular and horizontal gene transfer in Cryptosporidium parvum. Genome Biology 5:R88 PMID 15535864
- ↑ de Koning et al. (2000) Lateral gene transfer and metabolic adaptation in the human parasite Trichomonas vaginalis. Mol Biol Evol 17:1769-73
- ↑ Doolittle WF (1998) You are what you eat: a gene transfer ratchet could account for bacterial genes in eukaryotic nuclear genomes. Trends in Genetics 14:307-11 PMID 9724962
- ↑ Hall CS et al. (2005) Contribution of horizontal gene transfer to the evolution of Saccharomyces cerevisiae. Eukaryot Cell 4:1102-1115
- ↑ Dujon B et al. {2004) Genome evolution in yeasts. Nature 430:35-44 PMID 15229592
- ↑ Gray MW (1993) Origin and evolution of organelle genomes. Curr Opin Genet Dev 3:884-90 PMID 8118213
- Adams KL Palmer JD (2003) Evolution of mitochondrial gene content: gene loss and transfer to the nucleus. Molecular Phylogenetics and Evolution 29:380–95. PMID 14615181
- ↑ Shrestha K et al. (2001) Evidence for paclitaxel from three new endophytic fungi of Himalayan yew of Nepal. Planta Med 67:374-6 PMID 11458463
- ↑ Robertson HM et al. (1996) Reconstruction of the ancient 'mariners' of humans. Nature Genetics 12:360-361 PMID 8630486
- ↑ Kondo N et al. (2002) Genome fragment of Wolbachia endosymbiont transferred to X chromosome of host insect. Proc Natl Acad Sci USA 99:14280-5
- ↑ Adoutte A et al. (2000) The new animal phylogeny: reliability and implications. Proc Natl Acad Sci USA 97:4453-6 PMID 10781043
- Steenkamp ET et al. (2006) The protistan origins of animals and fungi. Mol Biol Evol 23:93-106 PMID 16151185
- ↑ Woese C et al. (1990) Towards a natural system of organisms: proposal for the domains Archaea, Bacteria, and Eucarya. Proc Natl Acad Sci USA 87 :4576-9 PMID 2112744
- Woese C, Fox G (1977). Phylogenetic structure of the prokaryotic domain: the primary kingdoms. Proc Natl Acad Sci USA 74:5088-90 PMID 270744
- ↑ Simonson AB et al. (2005) Decoding the genomic tree of life. Proc Natl Acad Sci USA 102 Suppl 1:6608-13 PMID 15851667
- Yap WH et al. (1999) Distinct types of rRNA operons exist in the genome of the actinomycete Thermomonspora chromogena and evidence for horizontal gene transfer of an entire rRNA operon. J Bacteriol 181:5201-9 PMID 10464188
- Gogarten JP Townsend JP (2005) Horizontal gene transfer, genome innovation and evolution. Nat Rev Microbiol 9:679-87 PMID 16138096
- ↑ Daubin V et al. (2003) Phylogenetics and the cohesion of bacterial genomes. Science 301:829-32 PMID 12907801
- Eisen JA, Fraser CM (2003) Viewpoint phylogenomics: intersection of evolution and genomics. Science 300:1706-7 PMID 12805538
- Fitzpatrick DA et al. (2006) A fungal phylogeny based on 42 complete genomes derived from supertree and combined gene analysis. BMC Evol Biol 6:99 PMID 17121679
- Ge F et al. (2005) The Cobweb of Life revealed by genome-scale estimates of horizontal gene transfer. PLoS Biology 3:e316 PMID 16122348
- Henz SR et al. (2005) Whole-genome prokaryotic phylogeny. Bioinformatics 21:2329-35 PMID 15166018
- Lerat E et al. (2005) Evolutionary origins of genomic repertoires in bacteria. PLoS Biology 3:e130 PMID 15799709
- Steenkamp ET et al. (2006) The protistan origins of animals and fungi. Mol Biol Evol 23:93-106 PMID 16151185]
- Urwin R, Maiden MC (2003) Multi-locus sequence typing: a tool for global epidemiology. Trends Microbiol 11:479-87 PMID 14557031
- ↑ Esser C et al. (2004) A genome phylogeny for mitochondria among alpha-proteobacteria and a predominantly eubacterial ancestry of yeast nuclear genes. Mol Biol Evol 21:1643-50 PMID 15155797
- Rivera MC, Lake JA (2004) The ring of life provides evidence for a genome fusion origin of eukaryotes. Nature 431:152-5 PMID 15356622
- Simonson AB et al. (2005) Decoding the genomic tree of life. Proc Natl Acad Sci USA 102 Suppl 1:6608-13 PMID 15851667
- ↑ Esser C et al. (2004) A genome phylogeny for mitochondria among alpha-proteobacteria and a preedominantly eubacterial ancestry of yeast nuclear genes. Mol Biol Evol 21:1643-50 PMID 15155797
- ↑ Kurland CG et al. (2006) Genomics and the irreducible nature of eukaryote cells. Science 312(5776):1011-4. PMID 16709776
Further Reading
- Horizontal Gene Transfer Syvanen M, Kado CI (2002) 2nd edition, Academic Press ISBN 0-12-680126-6 A comprehensive treatise. Reviewed here by M-W Ho
- Acquiring genomes: a theory of the origin of species. Margulis L and Sagan D (2002) Basic Books ISBN 0-465-04392-5. A book that looks at gene transfer from a different perspective to many conventional interpretations, but with an interesting emphasis on microbial diversity. Reviewed here.
- Where Do All Those Genes Come From? This study resolves a long-standing paradox: how is it possible to deduce reliable evolutionary histories from gene sequences in bacteria despite extensive HGT?
- Richardson AO, Palmer, JD (2007) Horizontal gene transfer in plants. Journal of Experimental Botany 58:1–9 doi:10.1093/jxb/erl148 PMID 17030541
- Gogarten JP Townsend JP (2005) Horizontal gene transfer, genome innovation and evolution. Nat Rev Microbiol 9:679-87 PMID 16138096. One article in a whole issue of the journal Nature Reviews Microbiology largely devoted to HGT.
- Mobile DNA. Berg DE, Howe MM (Eds.)(1989) American Society for Microbiology. Washington D.C. ISBN 1-55581-005-5 Book with a comprehensive discussion of mobile DNA, jumping genes, transposons etc in many organisms, not only bacteria. Reviewed by Freeling M (1990) in Q Rev Biol 65:217-8
- Salzberg SL et al. (2001) Microbial genes in the human genome: lateral transfer or gene loss? Science 292:1903-6 PMID 11358996. This reports that one dramatic claim of HGT - in which a distinguished group of scientists claimed that bacteria transferred their DNA directly into the human lineage - was simply wrong [1]
- Weinbauer MG, Rassoulzadegan F (2004) Are viruses driving microbial diversification and diversity? Envir Microbiol 6:1-11 PMID 14686936 Discussion of both the evolutionary and ecological activities of viruses in the ocean, a major source of HGT in nature.
- Woese C (2002) On the evolution of cells. Proc Natl Acad Sci USA 99:8742-7 PMID 12077305 This article shifts the emphasis in early phylogenic adaptation from vertical to horizontal gene transfer.
- Hall C et al. (2005) Contribution of horizontal gene transfer to the evolution of Saccharomyces cerevisiae. Eukaryot Cell 4:1102-15 PMID 15947202 Convincing evidence of horizontal transfer of bacterial DNA into yeast.
- Snel B et al. (1999) Genome phylogeny based on gene content. Nature Genetics 21:66-7 PMID 9916801 Proposal for using the presence or absence of a set of genes to infer phylogenies, in order to avoid confounding factors such as HGT.
External links
- Focus on horizontal gene transfer Webfocus in Nature with free access review articles
- Horizontal gene transfer among genomes: The complexity hypothesis. Proc Natl Acad Sci USA 96:3801-6 (1999)
- Lateral gene transfer and the nature of bacterial innovation (pdf), Ochman H et al (2000)
- Smallpox knows how to make a mouse protein. How did smallpox learn that? The New Yorker July 12, 1999, p44-61 'The poxviruses are promiscuous at capturing genes from their hosts,' Esposito said. 'It tells you that smallpox was once inside a mouse or some other small rodent'
- Gogarten Laboratory Webpages.
- Retrotransfer or gene capture: a feature of conjugative plasmids, with ecological and evolutionary significance
- Horizontal gene transfer - A new paradigm for biology
- Horizontal gene transfer (p334 of Molecular Genetics by Ulrich Melcher).
- Report on horizontal gene transfer by Mae-Wan Ho, 1999
- Recent evidence confirms risks of horizontal gene transfer
- Horizontal gene transfer at sciences.sdsu.edu